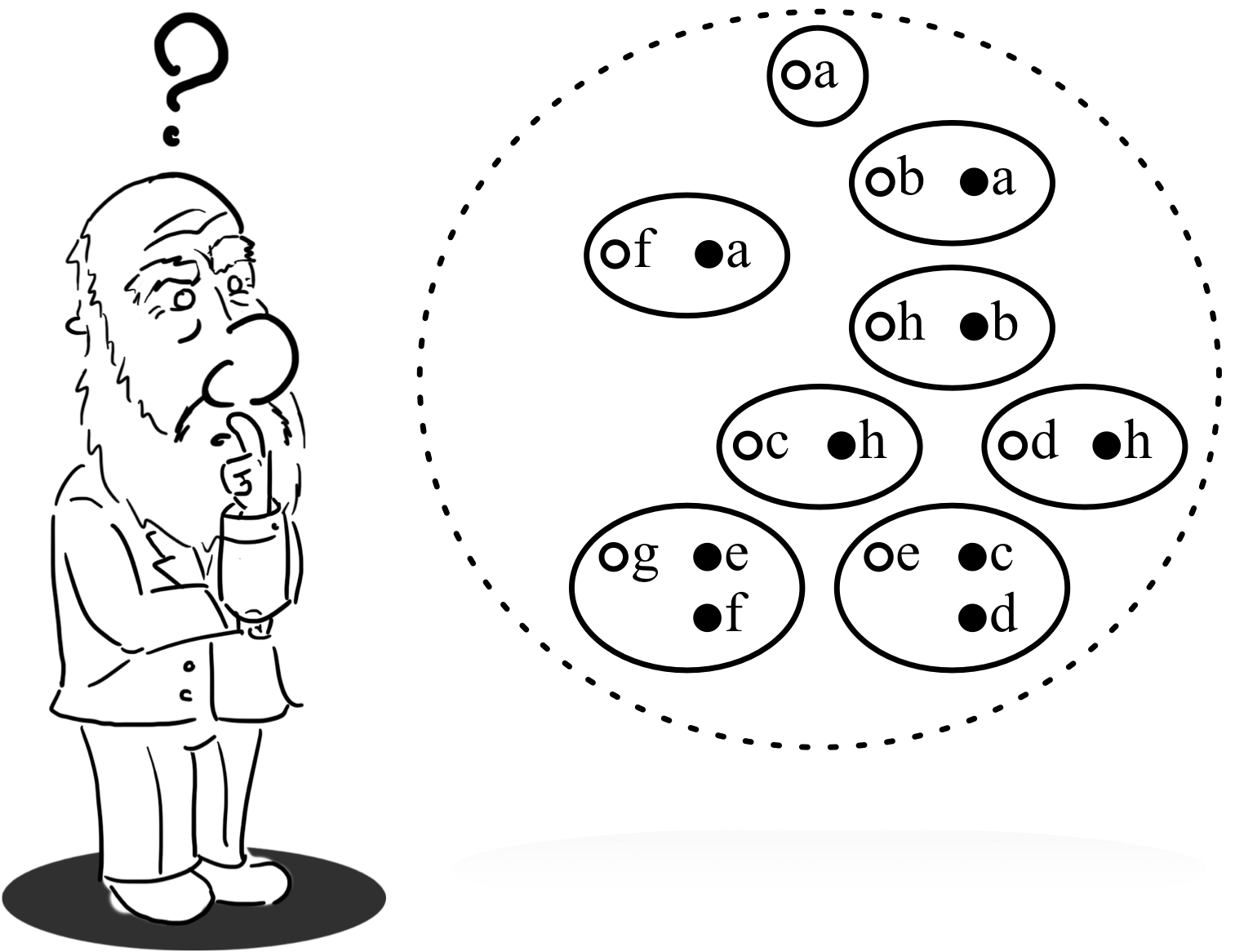
About DNs

Darwinian networks (DNs) were proposed to simplify working with Bayesian network (BNs). Rather than modelling the variables in a problem domain, DNs represent the probability tables in the model. The graphical manipulation of the tables then takes on a biological feel, where a conditional probability table (CPT) $P(X|Y)$ is viewed as the novel representation of a population $p(C,D)$ using both combative traits $C$ (coloured clear) and docile traits $D$ (coloured dark).
DNs can unify modeling and reasoning tasks into a single platform. DNs can represent exact inference using either variable elimination or arc-reversal, lazy propagation, as well as how DNs can represent testing independencies. Evolution is used to represent inference. The query $P(X|Y)$ posed to a BN ${\cal B}$ is represented by DN ${\cal D}^{\prime} = \{ p(X,Y) \}$. Adaptation is used to represent the testing of an independence $I_{\cal{B}}(X,Y,Z)$ holding in a BN $\cal{B}$. The test of independence $I_{\cal{B}}(X,Y,Z)$ holds in a BN $\cal{B}$ if and only if the adaptation $A({\cal{P}}_X, {\cal{P}}_Y, {\cal{P}}_Z)$ succeeds in the DN $\cal{D}$ for $\cal{B}$.